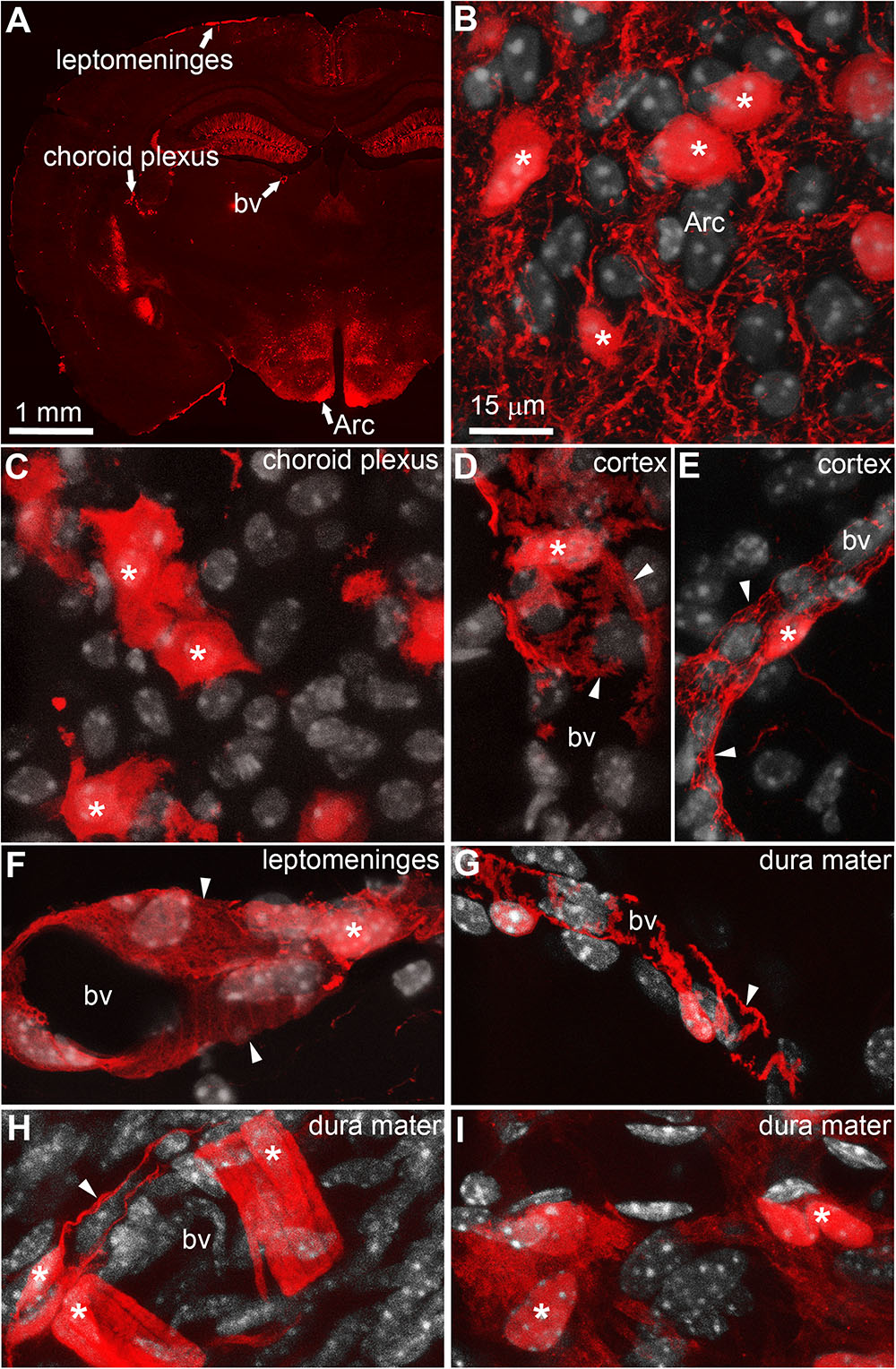
Upper panel: (b = 0) anatomical images (axial slice and coronal reconstruction) used for signal-to-noise ratio (SNR) estimation: ROI 1 (r = 20 voxels) is located in a region without signal in order to estimate the noise, ROI 2 (r = 5 mm) is located in the ventricles in order to estimate the signal intensity in a region with high (b = 0)-signal. 
cryogenic cooled resonator (CCR – right). Metrics applied for the evaluation of the quality of the resulting fiber data include the quality of the FA-maps and the quality of fiber reconstruction, which is tested quantitatively by tractwise fractional anisotropy statistics (TFAS). The aim is to keep the overall scanning time at approximately 30 minutes in order to enable the application of the scan protocol to in-vivo cohort studies. The SNR gain is supposed to generate high fidelity DTI data with high in-plane resolution and thin slices in order to get close to isotropic voxel resolution, which improves isotropic 3D DTI fiber reconstruction. Cryogenically cooled resonators (CCR) have been shown to significantly increase the effective SNR (factor between 2.4 and 2.5 in structural MRI scans) at 4.7T as well as at 9.4 T and are supposed to provide an SNR gain of at least a factor of 2 at 11.7T systems. Reduction of scanning time without compromising spatial resolution is obtained by application of a cryogenic coil at 11.7T field strength. The aim of the study was to investigate the feasibility of in vivo microstructural analysis of the mouse brain including fiber tracking by applying a rapid data acquisition protocol providing sufficient spatial resolution in short scan times. Although the spatial resolution revealed acceptable FT information, the corresponding long scanning time of about 90 minutes limits the application of the proposed protocol for large cohort studies. 
In-vivo experiments have been reported with in-plane resolution down to 156 µm×156 µm (matrix size 128×92) with an axial slice thickness of 375 µm and 500 µm. cannot be ensured by the in-vitro approach. Furthermore, the main advantage of DTI enabling a safe and non-invasive approach for longitudinally investigating normal development, aging, disease progression, etc. Obviously, the in-vitro scans cannot be applied to longitudinal studies. High-resolution experiments have been conducted on in-vitro brain tissue even to conduct fiber tracking (FT) and consequent tract based spatial statistics (TBSS). Total acquisition times for a slice thickness of 500 µm and lower were reached by acquisition times of 1–2 hours. Current in-plane resolutions for in-vivo experiments were ranging between 117 µm and 160 µm with a slice thickness between 375 µm and 1 mm. DTI studies of the rodent brain in high-field animal scanners with 7.0 up to 11.7T – require long scan times to ensure appropriate signal-to-noise ratio (SNR) at sufficient spatial resolution (see Supplementary Table S1). This allows for anatomical connectivity studies providing insights to brain function and morphology under physiological conditions or to dysfunction under pathological circumstances. Tractography algorithms use this information to track the neural pathways based on the assumption that the dominant direction of water motion – the main axis of the diffusion tensor – aligns with the fibers' orientations in an imaged voxel. DTI characterizes the diffusion of water molecules in tissue by using motion-probing spatial encoding : For each voxel of the image, the diffusion tensor describes the magnitude and directionality of the water movement (anisotropy). However, to establish DTI as a broad pre-clinical research tool to study white matter alterations in animal models, further optimization of the DTI protocols especially regarding acceptable data acquisition times appears mandatory.Īs an in-vivo approach, DTI is able to map the brain fiber directions and to render its 3D architecture, thus reconstructing axonal tracts and especially tracking the white matter pathways of the human brain. In the last decade, diffusion tensor imaging (DTI), has evolved as an increasingly important tool for studying the anatomy of the mouse brain in-vitro – as well as in-vivo –.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
